How Does DNA Regulate Cell Division? 1
How Does DNA Regulate Cell Division? 1
Luke Funk, Kuan-Chung Su, Iain Cheeseman, Paul Blainey
MIT Department of Biological Engineering, Koch Institute at MIT, Broad Institute
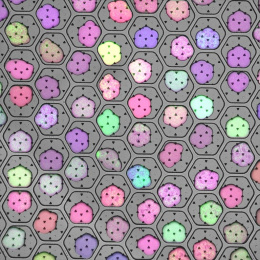
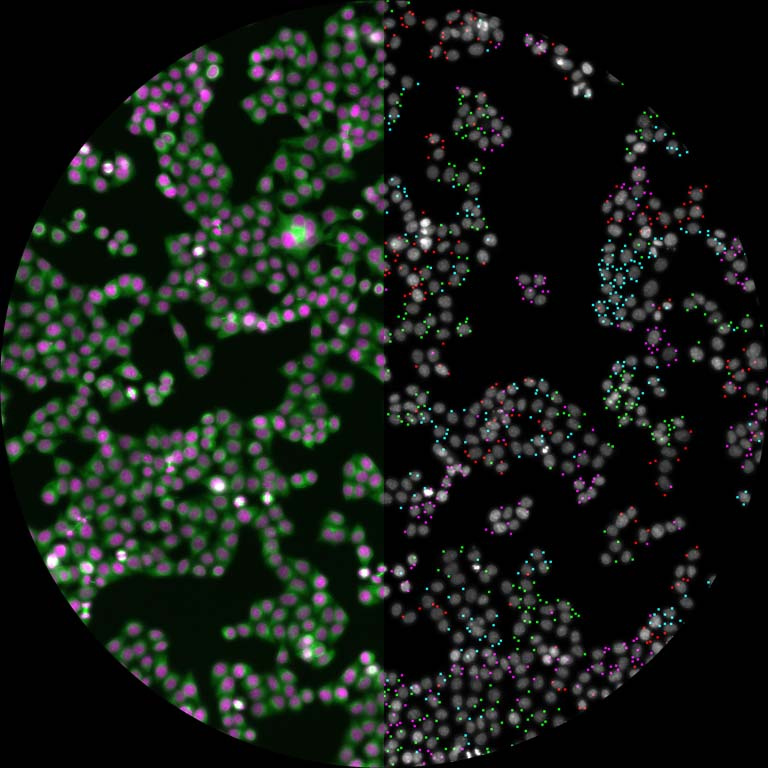
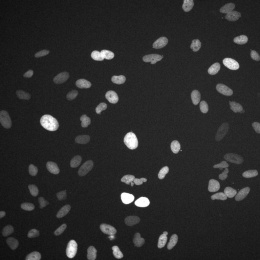
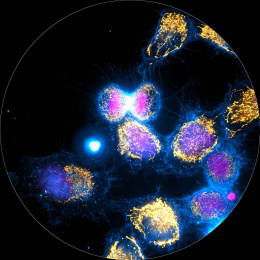
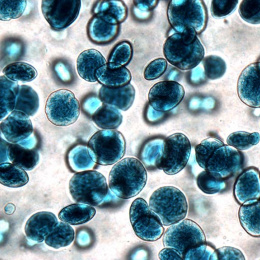
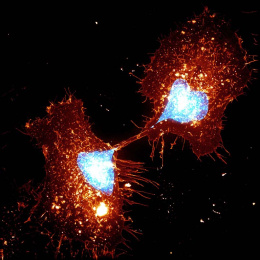
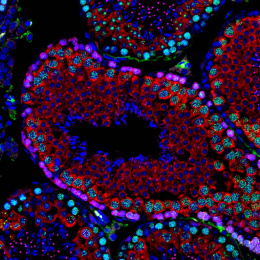
This is a split image showing two different views on the left and right of the same field of view from a fluorescence microscopy dataset. On the left are human cells stained to show their distribution of DNA (magenta color) as well as a protein known as tubulin (green color). Imaging these structures is helpful in identifying defects in mitosis (cell division), which is a fundamental process of life. In addition, each of the cells in this experiment has a different gene disrupted (“knocked-out”) using the CRISPR-Cas9 gene editing technology. The identity of which gene is knocked out in each cell is determined by reading a corresponding DNA barcode sequence that is expressed by each of the cells. On the right, you can see the location of these in situ DNA barcode reads, with the different colors representing one DNA base of the barcode sequence (green=“G”, red=“T”, magenta=“A”, cyan=“C”). This image-based “pooled” screening approach where cells with different gene knockouts are mixed and then the resulting phenotypes imaged is a new way to perform high throughput experiments to understand the functions of genes.
The goal of the experiment that all of these images came from was to profile thousands of genes known to be “essential” for the growth of human cell lines, looking for those genes that are involved in cell division or the regulation of mitosis. These processes go awry in a variety of human diseases, in particular many types of cancer. Better understanding the molecular players in these processes will hopefully enable increased understanding of oncogenesis and potentially identify new therapeutic targets.