Mapping the Body's Social Network 2
Mapping the Body's Social Network 2
Carly Ziegler, Shaina Carroll, Leslie Kean, Alex Shalek
Koch Institute at MIT, MIT Department of Chemistry, Institute of Medical Engineering and Science
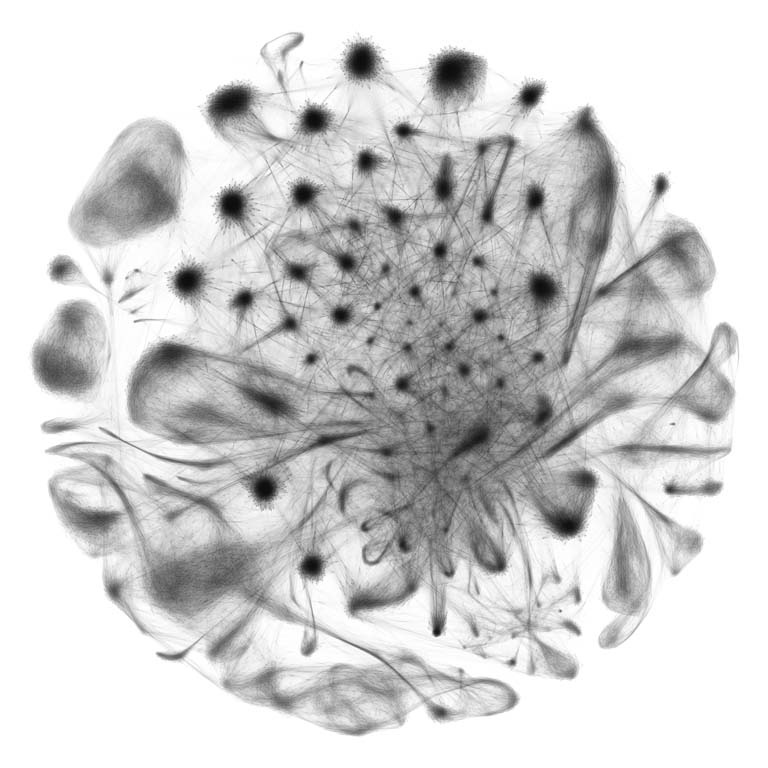
mRNA is the molecular language cells use to instruct changes to their appearance, function, and interaction with the world. Single-cell RNA-sequencing takes a snapshot of the total mRNA in an individual cell, providing insight into a cell’s current behavior, history, and trajectory. The Shalek lab (MIT – IMES, Chemistry, KI) has pioneered methods for ultra-high-throughput single-cell RNA-sequencing, enabling deep understanding of cellular physiology at an unprecedented scale. Here, we represent data comprising the first draft of a Non-Human Primate single cell atlas (45,782 cells, 4 animals, 14 tissues per animal). Single points on this graph represent single cells, their location and the lines between them reflect similarities in their mRNA content. Local areas of high density of points suggest cells of a common phenotype (e.g., activated microglia in the brain), and the point and edge colors reflect the tissue of origin. By characterizing a cell’s “type” purely by its transcriptome, we can validate and redraw cellular taxonomies. Collectively, this analysis allows us to begin to map the full complexity of cellular specialization and diversity in mammals.